Oligonucleotide therapeutics continue to reshape drug development by targeting disease pathways at the genetic level. Antisense oligonucleotides (ASOs), small interfering RNAs (siRNAs), and antibody-RNA conjugates (ARCs) are expanding the range of treatable diseases, particularly for targets that were previously considered inaccessible to conventional therapeutics.
As these modalities evolve, bioanalytical workflows face increasing pressure to deliver both sensitivity and selectivity across increasingly complex biological matrices. Oligonucleotide therapeutics present analytical challenges because they are too complex for standard LC-MS approaches and too heterogeneous for conventional ligand-binding assays.
Fit-for-purpose LC-MS strategies are increasingly important for oligonucleotide bioanalysis across plasma and tissue matrices.
Why Oligonucleotide Therapeutics Challenge Traditional Bioanalysis
Oligonucleotides occupy a unique analytical space between small molecules and biologics. Their structural complexity creates distinct challenges during quantitation, metabolite identification, and tissue analysis.
Ligand-binding assays and PCR-based approaches remain valuable for ultra-sensitive detection, particularly for high-potency therapeutics requiring low picogram-per-milliliter sensitivity. However, these methods can struggle to differentiate closely related metabolites generated through nuclease activity in vivo.
N-1 and N-2 metabolites frequently resemble the parent compound closely enough to bind the same capture probes, reducing assay specificity. Narrow dynamic ranges can also limit accurate quantitation across tissues where drug concentrations vary substantially.
Mass spectrometry-based workflows help address these limitations by improving molecular selectivity and enabling more robust characterization across plasma and tissue matrices.
Choosing the Right LC-MS Platform for Oligonucleotide Quantification
Different analytical questions require different LC-MS strategies. Successful oligonucleotide bioanalysis depends on selecting platforms that align with study objectives, matrix complexity, and sensitivity requirements.
Direct LC-MS/MS for Quantitative Bioanalysis
Triple quadrupole LC-MS/MS systems remain essential for regulated bioanalysis and toxicology studies. Multiple Reaction Monitoring (MRM) improves selectivity by separating parent oligonucleotides from structurally similar metabolites while supporting quantitation in the nanogram-per-milliliter range.
These systems are particularly effective when robust targeted quantitation is required across validated workflows.
High-Resolution Mass Spectrometry Improves Metabolite Characterization
High-resolution mass spectrometry (HRMS) provides enhanced resolving power for discovery-stage characterization and impurity profiling.
Phosphorothioate oligonucleotides commonly undergo desulfurization events that may co-elute with the parent compound during chromatographic separation. HRMS enables clearer visualization of isotopic envelopes impurities such as desulfurization that are difficult to resolve using direct LC-MS methods alone.
Hybridization LC-MS/MS Combines Sensitivity and Selectivity
Hybridization LC-MS/MS workflows integrate analyte enrichment with mass spectrometry detection. Capture probes selectively enrich oligonucleotides prior to LC-MS analysis, combining the sensitivity advantages of ligand-binding assays with the specificity of mass spectrometry.
This hybrid approach has become increasingly important for low-abundance oligonucleotide therapeutics in complex biological matrices.
Tissue Bioanalysis Requires Optimized Extraction Chemistry
Tissue analysis introduces a separate layer of complexity because oligonucleotides can bind intracellular proteins and become sequestered from detection.
Proteinase K digestion significantly improves analyte recovery by releasing bound oligonucleotides from tissue-associated proteins. Without digestion, recovery can decrease dramatically, limiting assay sensitivity and quantitative accuracy.
Digestion conditions also influence workflow efficiency and matrix cleanliness. Traditional Proteinase K digestion protocols operate overnight at 37°C, while thermostable alternatives can complete digestion in approximately one hour at elevated temperatures. Faster workflows can compress a two-day digestion workflow into a single day but may increase matrix interference and reduce signal-to-noise ratios.
Optimizing extraction conditions remains critical for achieving reproducible tissue bioanalysis.
Surrogate Tissue Matrices Can Streamline Early Discovery Workflows
Multi-organ bioanalysis often requires separate calibration matrices for liver, kidney, heart, spleen, and additional tissues. Maintaining individual matrix calibrators increases operational complexity and cost.
Surrogate tissue matrices offer an alternative strategy for discovery-stage studies. By combining multiple tissue types into a pooled matrix, laboratories can simplify workflow development while maintaining comparable analytical performance across organs.
This approach may simplify workflow development for discovery-stage tissue assessments.
The 1.5-Plex Workflow Conserves Sample Volume in ARC Bioanalysis
Antibody-RNA conjugates introduce additional analytical complexity because both the antibody carrier and RNA payload require characterization.
Running separate assays for total antibody and conjugated RNA increases sample consumption, creating challenges in pediatric studies and low-volume preclinical models.
The 1.5-plex workflow addresses this limitation through a single immunocapture followed by split analytical processing. One aliquot undergoes base hydrolysis for RNA payload quantitation, while the second aliquot undergoes tryptic digestion for surrogate peptide analysis of total antibody.
Careful calibration design is equally important. Total conjugate concentration does not necessarily reflect RNA payload concentration, making analyte-specific stock definitions essential for accurate quantitation.
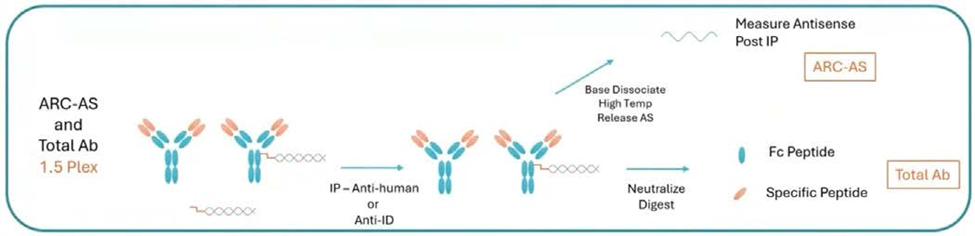
(1.5-plex LC-MS workflow for antibody-RNA conjugate bioanalysis)
Instrument Selection Should Be Driven by Matrix Performance
Higher instrument sensitivity did not universally improve assay performance across all matrices evaluated.
Matrix effects can significantly alter signal-to-noise ratios depending on tissue type and assay conditions. In some plasma applications, higher ion sampling may also increase background noise, reducing overall assay clarity. Conversely, more sensitive platforms may become essential for achieving reportable detection limits in difficult tissue matrices such as heart tissue.
Fit-for-purpose platform selection should therefore prioritize real sample performance rather than relying solely on instrument specifications.
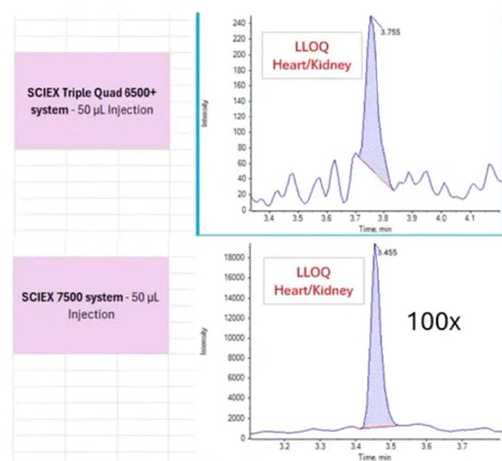
(Comparison of LC-MS sensitivity in oligonucleotide tissue bioanalysis)
Operational Controls Influence Long-Term LC-MS Robustness
Long-term oligonucleotide bioanalysis performance depends heavily on operational consistency and laboratory controls.
Dedicated LC-MS Systems Reduce Cross-Contamination Risk
Ion-pairing reagents such as triethylamine (TEA) and N,N-diisopropylethylamine (DIPEA) can persist on LC system surfaces and suppress ionization in unrelated assays. Dedicated instrumentation helps minimize cross-contamination and ion suppression in unrelated assays.
Surface Passivation Improves Recovery Consistency
Oligonucleotides readily adsorb to standard polypropylene consumables, particularly in low-protein tissue homogenates. DNA and Protein LoBind labware help reduce sample loss and improve recovery reproducibility across calibration ranges.
Mobile Phase Chemistry Directly Impacts Sensitivity
Minor changes in ion-pairing chemistry can substantially alter assay performance. Replacing TEA with DIPEA may improve charge masking, peak sharpness, and desolvation efficiency, leading to improved analytical sensitivity.
Mobile phase optimization remains one of the most impactful variables in oligonucleotide LC-MS method development.
The Future of Oligonucleotide Bioanalysis Requires Integrated LC-MS Strategies
As oligonucleotide therapeutics continue to expand across clinical pipelines, bioanalytical methods must evolve to support increasingly complex modalities and matrices.
No single platform or workflow solves every challenge in oligonucleotide bioanalysis. Successful strategies integrate selective mass spectrometry platforms, optimized extraction chemistry, matrix-aware calibration approaches, and operational controls that maintain long-term robustness.
Fit-for-purpose LC-MS workflows will remain central to advancing reliable oligonucleotide quantitation from plasma to tissue.
Watch the Full Presentation
To explore the complete discussion on oligonucleotide bioanalysis strategies, including LC-MS platform selection, tissue extraction workflows, and operational considerations for plasma and tissue assays, watch the full presentation featuring Rathna Veeramachaneni from KCAS Bio. The session provides additional scientific context and practical examples supporting fit-for-purpose oligonucleotide bioanalysis workflows. This webinar was a collaboration between Sciex and KCAS Bio.

